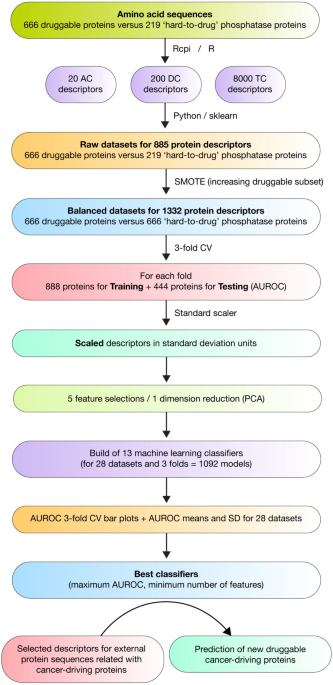
Overview of the Study
This research focuses on developing advanced machine learning models to predict proteins that can be targeted for cancer treatment. By analyzing protein sequences, the study identifies which proteins are likely to be druggable. It employs three types of amino acid composition descriptors: amino acid composition (AC), di-amino acid composition (DC), and tri-amino acid composition (TC). These descriptors are crucial for understanding the structure and function of proteins, which in turn helps in predicting their interactions with drugs.
Key Findings
- The study successfully predicted 2,080 out of 2,339 cancer-driving proteins as druggable, showing a high accuracy rate.
- The best-performing model achieved an area under the receiver operating characteristic (AUROC) score of 0.992, indicating strong predictive power.
- Selected features from the analysis included significant amino acid patterns that are biologically relevant, such as HME and NSH.
- The research identified 23 key druggable proteins with unfavorable prognostic significance, highlighting their potential as therapeutic targets.
Importance of the Research
Understanding which proteins are druggable is essential for developing new cancer therapies. This study’s findings can streamline the drug discovery process by providing a list of promising targets. The innovative machine learning approach not only enhances the accuracy of predictions but also contributes to personalized medicine by tailoring treatments based on genetic profiles. The insights gained from this research will significantly impact future cancer treatment strategies and drug development efforts.